The “dot plots in R” typically refers to two distinct types of visualizations: (i) Cleveland dot plots (for comparing categories) and (ii) Stacked (Wilkinson) dot plots (for showing distributions). A dot plot is a graphical representation that breaks the range of data into many small equal-width intervals and counts the number of observations in each interval. The interval count is superimposed on the number line at the interval midpoint as a series of dots (stacked if repeated), usually one for each observation. For $mpg$ from the $mtcars$ dataset, the intervals are centered at integer values, so the display gives the number of observations at each distinct observed head breadth.
Table of Contents
Dot Plots in R Language using Base and ggplot2 Packages
Plotting Dot Plot using R Base Graphics
The following code may be used to draw a dot plot using R Base Graphics
attach(mtcars)
par(mfrow = c(3, 1))
# Dot Plot 1
stripchart(mpg, main = "Miles per Gallon", xlab = "mpg")
# Dot Plot 2
stripchart(mpg, method = "stack", cex = 2,
main = "Miles Per Gallon (with Stack Method)")
# Dot Plot 3
stripchart(mpg, method = "jitter", cex = 2, frame.plot = FALSE,
main = "Mile Per Gallon (with no frame & Jitter Method")Plotting a Dot Plot using the ggplot Package
The following code may be used to draw dot plots in R using the ggplot2 package:
library(ggplot2)
library(gridExtra)
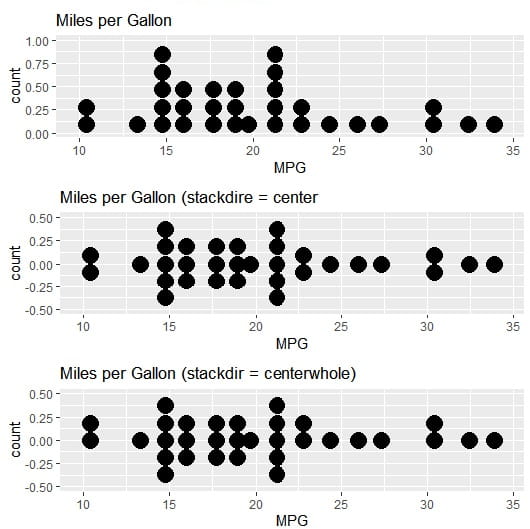
# Dot Plot 1
p1 <- ggplot(mpg, aes(x = mpg))
p1 <- p1 + geom_dotplot(binwidth = 2)
p1 <- p1 + labs(title = "Miles per Gallon")
p1 <- p1 + xlab("MPG")
# Dot Plot 2
p2 <- ggplot(mpg, aes(x = mpg))
p2 <- p2 + geom_dotplot(binwidth = 2, stackdir = "center")
p2 <- p2 + labs(title = "Miles per Gallon (stackdire = center")
p2 <- p2 + xlab("MPG")
# Dot Plot 3
p3 <- ggplot(mpg, aex(x = mpg))
p3 <- p3 + geom_dotplot(binwidth = 2, stackdir = "centerwhole")
p3 <- p3 + labs(title = "Miles per Gallon (stackdir = centerwhole)")
p3 <- p3 + xlab("MPG")
grid.arrange(grobs = list(p1, p2, p3), ncol =1)Adjust Binwidth: You can manually set the binwidth parameter to change the size of the bins the dots fall into. This helps adjust the granularity of the visualization.
Dot Plots in R Group by a Categorical Variable
One can use a categorical variable, such as cyl (number of cylinders), to group the dots and display the distribution for each group. The cyl variable needs to be converted to a factor first for proper display. The code is
ggplot(mtcars, aes(x = mpg, fill = factor(cyl))) + # Map 'cyl' to fill aesthetic geom_dotplot(binwidth = 1.5) + labs(fill = "Cylinders") # Add a label to the legend
Key Advantages of Dot Plots in R
- Transparency: They show raw data, revealing gaps, clusters, and outliers that summary plots obscure.
- Small Sample Size Clarity: Unlike boxplots, they don’t hide sample size or become misleading with n < 10.
- Quantitative Comparisons: Cleveland dot plots are superior to bar charts for comparing many categories because they use position (not bar length), reducing visual clutter.
- Flexibility: With R packages (
ggplot2,ggbeeswarm,ggdist), you can layer uncertainty intervals, trend lines, and faceting to handle complex datasets.
Frequently Asked Questions about Dot Plots in R
How to add mean and median lines to a dot plot in R
First, we need summary statistics (stat_summary()) that will be overlaid on individual points.
ggplot(iris, aes(x = Species, y = Sepal.Length)) +
geom_dotplot(binaxis = "y", stackdir = "center",
dotsize = 0.6, alpha = 0.5) +
stat_summary(fun = mean, geom = "point",
shape = 18, size = 4, color = "red") +
stat_summary(fun = median, geom = "point",
shape = 15, size = 3, color = "blue") +
labs(title = "Red = Mean, Blue = Median")How to create a Horizontal (Cleveland) Dot Plot in R?
One can compare many categories where vertical space is limited. One needs to swap the $x$ and $y$ axes and use coord_flip() to flip the horizontal geometry.
# Method 1: Flip coordinates ggplot(mtcars, aes(x = reorder(rownames(mtcars), mpg), y = mpg)) + geom_point(size = 2) + coord_flip() + labs(x = "Car Model", y = "MPG", title = "Car Fuel Efficiency Ranking") # Method 2: Direct horizontal with reorder ggplot(mtcars, aes(x = mpg, y = reorder(rownames(mtcars), mpg))) + geom_point(size = 2, color = "steelblue") + labs(x = "MPG", y = "", title = "Horizontal Dot Plot")
How to color dots by a third variable?
One can use an additional dimension (such as color by transmission type) and map a variable to fill or color aesthetic.
# Color by transmission (am = 0 automatic, 1 manual)
ggplot(mtcars, aes(x = factor(cyl), y = mpg, fill = factor(am))) +
geom_dotplot(binaxis = "y", stackdir = "center",
dotsize = 0.7, alpha = 0.7) +
scale_fill_manual(values = c("lightblue", "orange"),
labels = c("Automatic", "Manual")) +
labs(fill = "Transmission")How to handle missing data in Dot plots?
NA values cause errors or gaps in the plot. One can remove NAs or handle missing values explicitly.
# Check for missing values sum(is.na(airquality$Ozone)) # Option 1: Remove NAs airquality_clean <- na.omit(airquality) ggplot(airquality_clean, aes(x = factor(Month), y = Ozone)) + geom_dotplot(binaxis = "y", stackdir = "center") # Option 2: Use na.rm in geom ggplot(airquality, aes(x = factor(Month), y = Ozone)) + geom_dotplot(binaxis = "y", stackdir = "center", na.rm = TRUE)
How to add a boxplot behind a dot plot?
Suppose you want to show both the distribution summary and raw data. The layer geom_boxplot() first and then geom_dotplot() with transparency.
ggplot(iris, aes(x = Species, y = Sepal.Length)) +
geom_boxplot(width = 0.3, alpha = 0.5, outlier.shape = NA) +
geom_dotplot(binaxis = "y", stackdir = "center",
dotsize = 0.5, alpha = 0.6, fill = "steelblue") +
labs(title = "Boxplot + Dot Plot Combination")How to adjust the dot size and spacing in dot plots?
Dots are too big or too small; one can adjust them using dotsize, binwidth, and stackratio.
# Control dot appearance
ggplot(mtcars, aes(x = factor(cyl), y = mpg)) +
geom_dotplot(binaxis = "y",
stackdir = "center",
dotsize = 0.4, # Dot size (smaller = less overlap)
binwidth = 1.5, # Controls grouping sensitivity
stackratio = 0.8) # Space between stacked dotsHow to create faceted dot plots (multiple panels)?
To compare subgroups across categories, use facet_wrap() or facet_grid().
# Facet by transmission type ggplot(mtcars, aes(x = factor(cyl), y = mpg)) + geom_dotplot(binaxis = "y", stackdir = "center", dotsize = 0.6) + facet_wrap(~ am, labeller = labeller(am = c(`0` = "Automatic", `1` = "Manual"))) + labs(title = "MPG Distribution by Cylinders and Transmission")