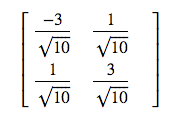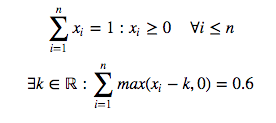I am learning SVD by following this MIT course.
the Matrix is constructed as
C = np.matrix([[5,5],[-1,7]])
C
matrix([[ 5, 5],
[-1, 7]])
the lecturer gives the V as
this is close to
w, v = np.linalg.eig(C.T*C)
matrix([[-0.9486833 , -0.31622777],
[ 0.31622777, -0.9486833 ]])
but np.linalg.svd(C) gives a different output
u, s, vh = np.linalg.svd(C)
vh
matrix([[ 0.31622777, 0.9486833 ],
[ 0.9486833 , -0.31622777]])
it seems the vh exchange the elements in the V vector, is it acceptable?
did I do and understand this correctly?
Solution:
For linalg.eig your Eigenvalues are stored in w. These are:
>>> w
array([20., 80.])
For your singular value decomposition you can get your Eigenvalues by squaring your singular values (C has maximum rank so everything is easy here):
>>> s**2
array([80., 20.])
As you can see their order is flipped.
From the linalg.eig documentation:
The eigenvalues are not necessarily ordered
From the linalg.svd documentation:
Vector(s) with the singular values, within each vector sorted in descending order. …
In general routines that give you Eigenvalues and Eigenvectors do not “sort” them necessarily the way you might want them. So it is always important to make sure you have the Eigenvector for the Eigenvalue you want. If you need them sorted (e.g. by Eigenvalue magnitude) you can always do this yourself (see here: sort eigenvalues and associated eigenvectors after using numpy.linalg.eig in python).
Finally note that the rows in vh contain the Eigenvectors, whereas in v it’s the columns.
So that means that e.g.:
>>> v[:,0].flatten()
matrix([[-0.9486833 , 0.31622777]])
>>> vh[:,1].flatten()
matrix([[ 0.9486833 , -0.31622777]])
give you both the Eigenvector for the Eigenvalue 20.