I am trying to understand the differences between the discrete convolution provided by Scipy and the analytic result one would obtain. My question is how does the time axis of the input signal and the response function relate the the time axis of the output of a discrete convolution?
To try and answer this question I considered an example with an analytic result. My input signal is a Gaussian and my response function is a exponential decay with a step function. The analytic results of the convolution of these two signals is a modified Gaussian (https://en.wikipedia.org/wiki/Exponentially_modified_Gaussian_distribution). 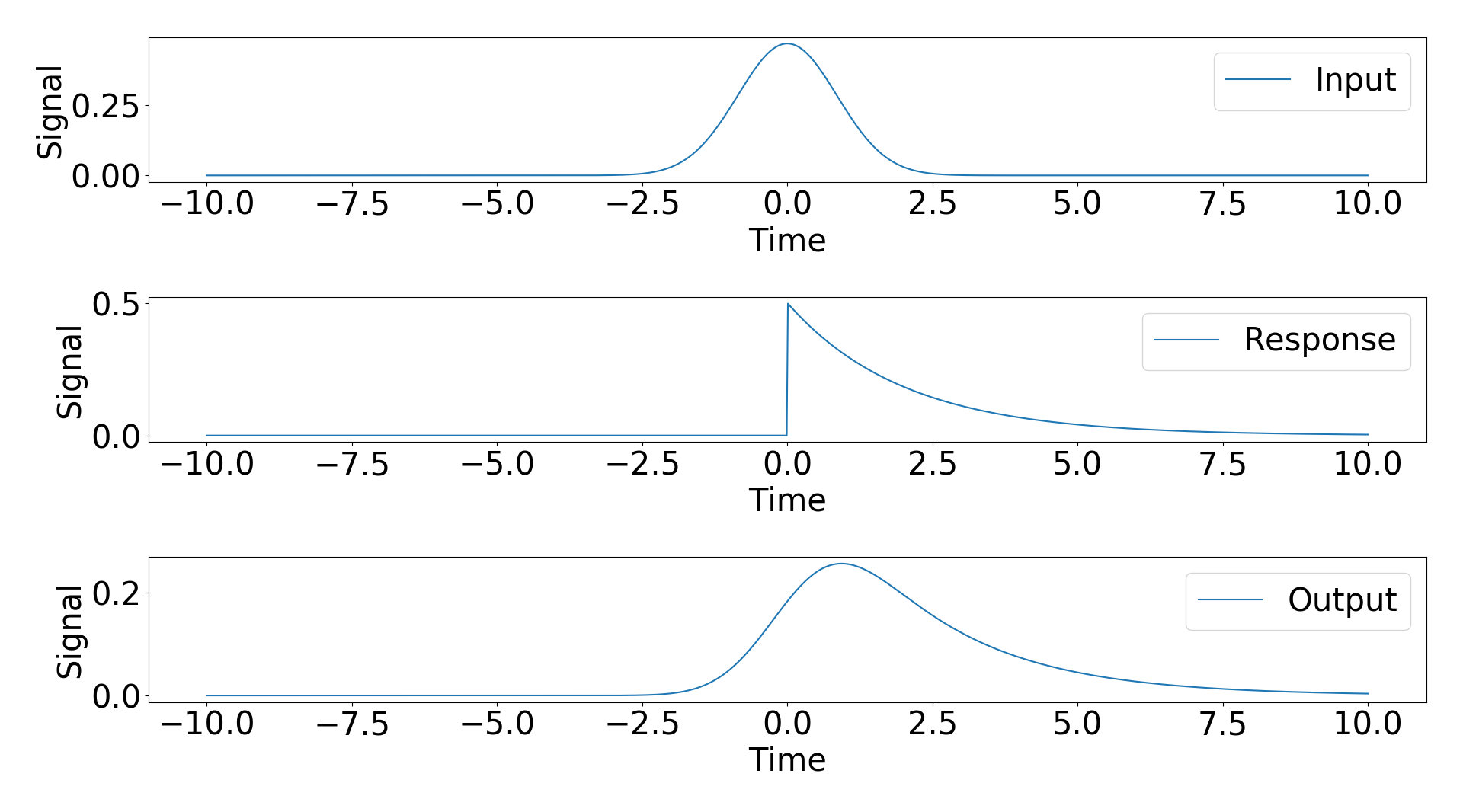
Scipy gives three modes of convolution, “same”, “full”, “valid”. I applied the “same” and “full” convolutions and plotted the result against the analyitic solution below.
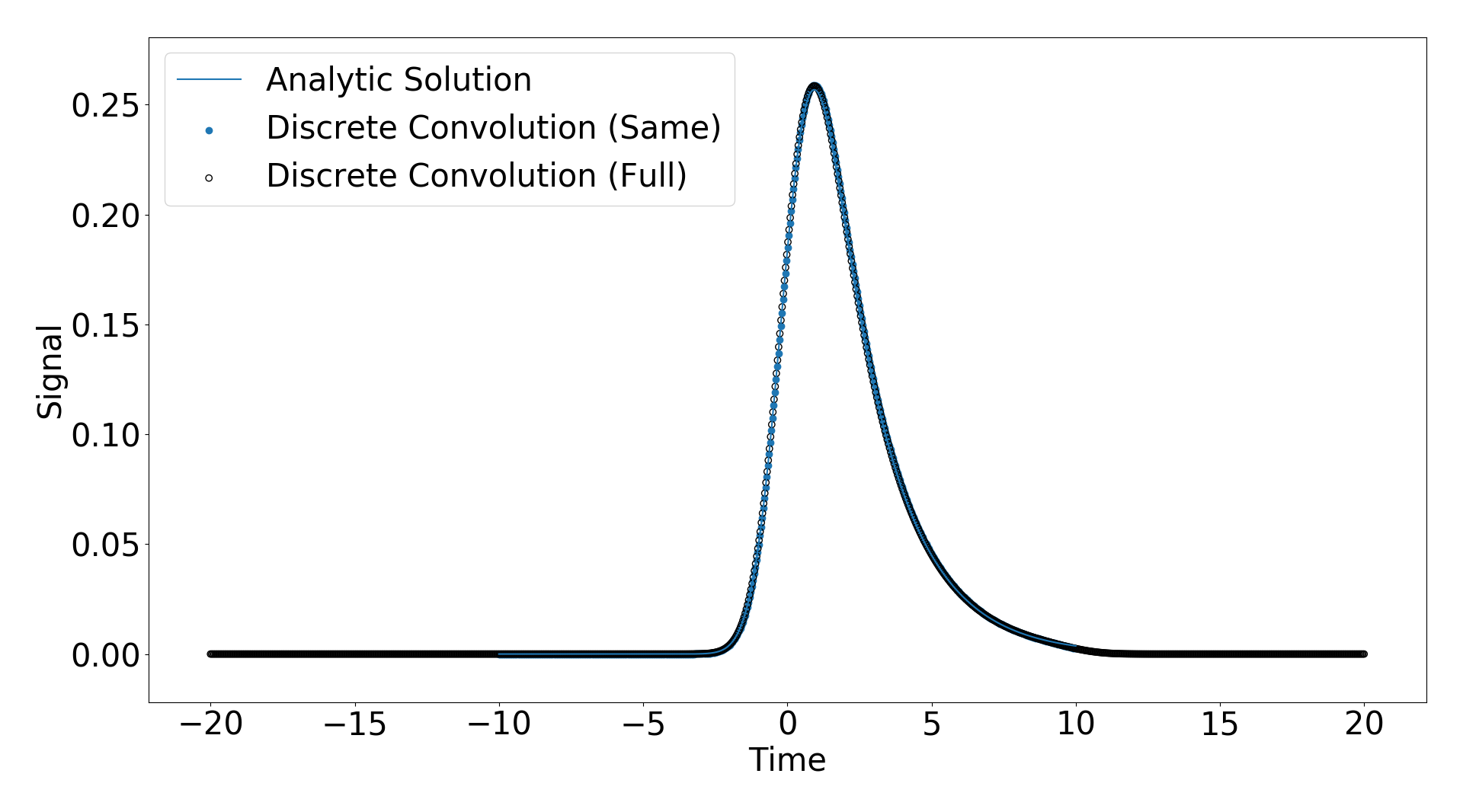
You can see they match extremely well.
One important feature to note is that for the “full” discrete convolution, the resulting time domain is much larger than the input signal time domain (see. https://www.researchgate.net/post/How_can_I_get_the_convolution_of_two_signals_in_time_domain_by_just_having_the_values_of_amplitude_and_time_using_Matlab), but for the “same” discrete convolution the time domains are the same and the input and response domains (which I have chosen to be the same for this example).
My confusion arose when I observed that changing the domain in which my response function was populated changed the result of the convolution functions. That means when I set t_response = np.linspace(-5,10,1000) instead of t_response = np.linspace(-10,10,1000) I got different results as shown below
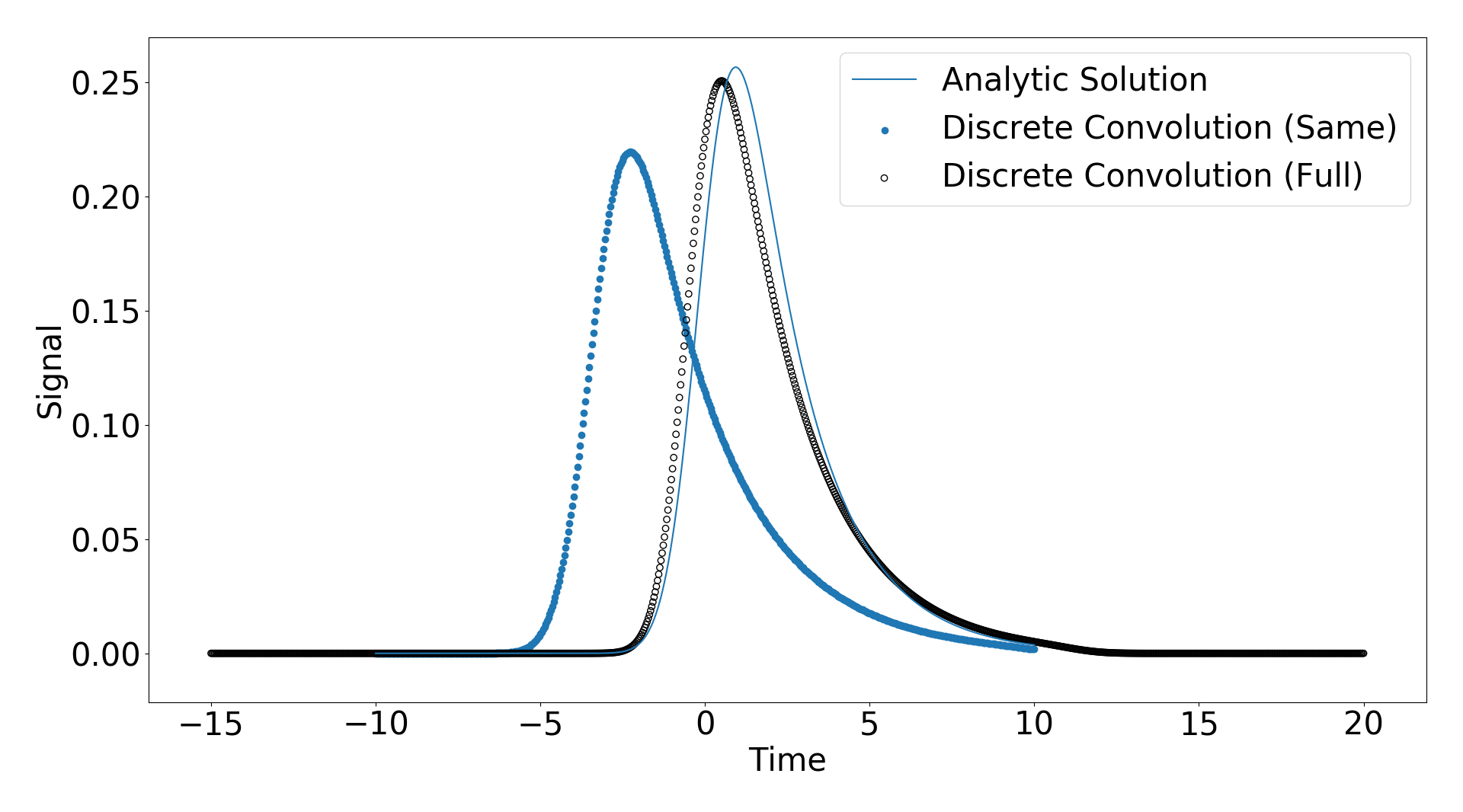
As you can see the solutions shift slightly. My question is how does the time axis of the input signal and the response function relate the the time axis of the output? I have attached the code I am using below and any help would be appreciated.
import numpy as np
import matplotlib as mpl
from scipy.special import erf
import matplotlib.pyplot as plt
from scipy.signal import convolve as convolve
params = {'axes.labelsize': 30,'axes.titlesize':30, 'font.size': 30, 'legend.fontsize': 30, 'xtick.labelsize': 30, 'ytick.labelsize': 30}
mpl.rcParams.update(params)
def Gaussian(t,A,mu,sigma):
return A/np.sqrt(2*np.pi*sigma**2)*np.exp(-(t-mu)**2/(2.*sigma**2))
def Decay(t,tau,t0):
''' Decay expoential and step function '''
return 1./tau*np.exp(-t/tau) * 0.5*(np.sign(t-t0)+1.0)
def ModifiedGaussian(t,A,mu,sigma,tau):
''' Modified Gaussian function, meaning the result of convolving a gaussian with an expoential decay that starts at t=0'''
x = 1./(2.*tau) * np.exp(.5*(sigma/tau)**2) * np.exp(- (t-mu)/tau)
s = A*x*( 1. + erf( (t-mu-sigma**2/tau)/np.sqrt(2*sigma**2) ) )
return s
### Input signal, response function, analytic solution
A,mu,sigma,tau,t0 = 1,0,2/2.344,2,0 # Choose some parameters for decay and gaussian
t = np.linspace(-10,10,1000) # Time domain of signal
t_response = np.linspace(-5,10,1000)# Time domain of response function
### Populate input, response, and analyitic results
s = Gaussian(t,A,mu,sigma)
r = Decay(t_response,tau,t0)
m = ModifiedGaussian(t,A,mu,sigma,tau)
### Convolve
m_full = convolve(s,r,mode='full')
m_same = convolve(s,r,mode='same')
# m_valid = convolve(s,r,mode='valid')
### Define time of convolved data
t_full = np.linspace(t[0]+t_response[0],t[-1]+t_response[-1],len(m_full))
t_same = t
# t_valid = t
### Normalize the discrete convolutions
m_full /= np.trapz(m_full,x=t_full)
m_same /= np.trapz(m_same,x=t_same)
# m_valid /= np.trapz(m_valid,x=t_valid)
### Plot the input, repsonse function, and analytic result
f1,(ax1,ax2,ax3) = plt.subplots(nrows=3,ncols=1,num='Analytic')
ax1.plot(t,s,label='Input'),ax1.set_xlabel('Time'),ax1.set_ylabel('Signal'),ax1.legend()
ax2.plot(t_response,r,label='Response'),ax2.set_xlabel('Time'),ax2.set_ylabel('Signal'),ax2.legend()
ax3.plot(t_response,m,label='Output'),ax3.set_xlabel('Time'),ax3.set_ylabel('Signal'),ax3.legend()
### Plot the discrete convolution agains analytic
f2,ax4 = plt.subplots(nrows=1)
ax4.scatter(t_same[::2],m_same[::2],label='Discrete Convolution (Same)')
ax4.scatter(t_full[::2],m_full[::2],label='Discrete Convolution (Full)',facecolors='none',edgecolors='k')
# ax4.scatter(t_valid[::2],m_valid[::2],label='Discrete Convolution (Valid)',facecolors='none',edgecolors='r')
ax4.plot(t,m,label='Analytic Solution'),ax4.set_xlabel('Time'),ax4.set_ylabel('Signal'),ax4.legend()
plt.show()
Solution:
The crux of the problem is that in the first case your signals have the same sampling rates, while in the second they do not.
I feel that it’s easier to think about this in the frequency domain, where convolution is multiplication. When you create a signal and filter with the same time axis, np.linspace(-10, 10, 1000), they now have the same sampling rates. Applying the Fourier transform to each, the resulting arrays give the power at the same frequencies for the signal and filter. So you can directly multiply corresponding elements of those arrays.
But when you create a signal with time axis np.linspace(-10, 10, 1000) and a filter with time axis np.linspace(-5, 10, 1000), that’s no longer true. Applying the Fourier transform and multiplying corresponding elements is no longer correct, because the frequencies at the corresponding elements is not the same.
Let’s make this concrete, using your example. The frequency of the first element of the transform (i.e., np.fft.fftfreq(1000, np.diff(t).mean())[1]) of the signal (s) is about 0.05 Hz. But for the filter (r), the frequency of the first element is about 0.066 Hz. So multiplying those two elements is multiplying the power at two different frequencies. (This subtlety is why you often see signal processing examples first defining the sampling rate, and then creating time arrays, signals, and filters based on that.)
You can verify that this is correct by creating a filter which extends over the time range you’re interested in [-5, 10), but with the same sampling rate as the original signal. So using:
t = np.linspace(-10, 10, 1000)
t_response = t[t > -5.0]
generates a signal and filter over different time ranges but at the same sampling rate, so the convolution should be correct. This also means you need to modify how each array is plotted. The code should be:
ax4.scatter(t_response[::2], m_same[125:-125:2], label='Same') # Conv extends beyond by ((N - M) / 2) == 250 / 2 == 125 on each side
ax4.scatter(t_full[::2], m_full[::2], label='Full')
ax4.scatter(t_response, m, label='Analytic solution')
This generates the plot below, in which the analytic, full, and same convolutions match up well.




