-
Notifications
You must be signed in to change notification settings - Fork 219
Description
Prework
- Read and agree to the code of conduct and contributing guidelines.
- If there is already a relevant issue, whether open or closed, comment on the existing thread instead of posting a new issue.
Description
The docs of gt::text_transform() say that the locations argument can be any of cells_body(), cells_stub(), cells_row_groups(), cells_column_labels(), and cells_column_spanners().
I am assuming that I may be abusing the function somewhat by wanting to apply html there. However, the behavior here is somewhat inconsistent. Depending on the value of locations, the applied function needs to output different things in order to work. I create three different function that all aim to bold the input text. I know there are other ways to bold column labels, this is a simplified example of what I am actually trying to do.
Is there a way to consistently apply html to all locations allowed in gt::text_transform()? That's especially a problem if a user applies multiple locations wrapped in a list.
Reproducible example
We create three different helper functions. The first one is simple, just paste text in-between bold tags. The second one applies gt::html but inside apply because htmltools::HTML collapses the input vector. The third one allows exactly this collapse.
# EASY STRING MANIPULATION. WORKS IN CELL BODIES
make_bold_1 <- function(text){
paste0("<b>", text, "</b>")
}
# SPECIFICALLY MAKE IT A HTML STRING
# NEEDS LAPPLY BECAUSE htmltools::HTML COLLAPSES THE VECTOR
make_bold_2 <- function(text){
lapply(paste0("<b>", text, "</b>"), gt::html)
}
# SPECIFICALLY MAKE IT A HTML STRING
# ALLOW htmltools::HTML TO COLLAPSE
make_bold_3 <- function(text){
gt::html(paste0("<b>", text, "</b>"))
}
example_data <- head(mtcars, 5)
gt_tbl <- gt::gt(example_data, groupname_col = "cyl")
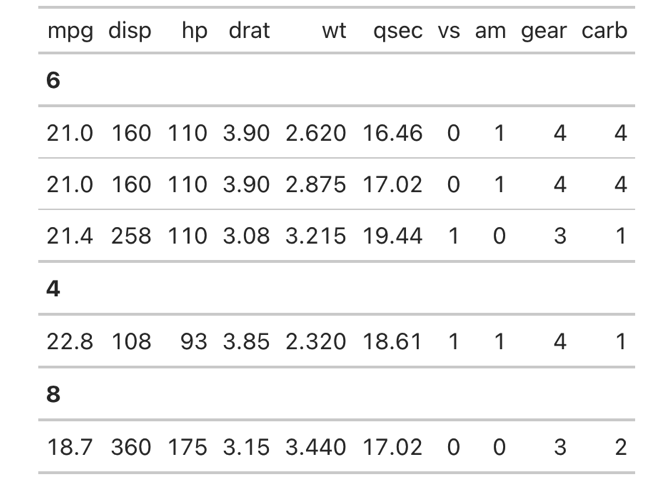
# OUR EXAMPLE TABLE WITHOUT ANY MODIFICATION
nflplotR::gt_render_image(gt_tbl)MODIFY CELL BODY
# SIMPLE MAKE BOLD WORKS IN CELL BODY
gt_tbl |>
gt::text_transform(make_bold_1, locations = gt::cells_body()) |>
nflplotR::gt_render_image()MODIFY ROW GROUP LABELS
# ... BUT NOT IN GROUPS
gt_tbl |>
gt::text_transform(make_bold_1, locations = gt::cells_row_groups()) |>
nflplotR::gt_render_image()# ... BUT THE LAPPLY VERSION WORKS
gt_tbl |>
gt::text_transform(make_bold_2, locations = gt::cells_row_groups()) |>
nflplotR::gt_render_image()# ... COLLAPSING MESSES IT UP
gt_tbl |>
gt::text_transform(make_bold_3, locations = gt::cells_row_groups()) |>
nflplotR::gt_render_image()MODIFY COLUMN LABELS
Neither of the 3 functions work.
gt_tbl |>
gt::text_transform(make_bold_1, locations = gt::cells_column_labels()) |>
nflplotR::gt_render_image()gt_tbl |>
gt::text_transform(make_bold_2, locations = gt::cells_column_labels()) |>
nflplotR::gt_render_image()gt_tbl |>
gt::text_transform(make_bold_3, locations = gt::cells_column_labels()) |>
nflplotR::gt_render_image()SWITCH TO gt::cols_label_with
gt_tbl |>
gt::cols_label_with(fn = make_bold_1) |>
nflplotR::gt_render_image()# THE LIST VERSION ERRORS BECAUSE OF WRONG OUTPUT TYPE
gt_tbl |>
gt::cols_label_with(fn = make_bold_2) |>
nflplotR::gt_render_image()
#> Error in `gt::cols_label_with()`:
#> ! `fn` must return a character vector.
#> Backtrace:
#> ▆
#> 1. ├─nflplotR::gt_render_image(gt::cols_label_with(gt_tbl, fn = make_bold_2))
#> 2. └─gt::cols_label_with(gt_tbl, fn = make_bold_2)
#> 3. └─cli::cli_abort("{.arg fn} must return a character vector.")
#> 4. └─rlang::abort(...)THE COLLAPSE VERSION WORKS?
gt_tbl |>
gt::cols_label_with(fn = make_bold_3) |>
nflplotR::gt_render_image()Created on 2023-09-06 with reprex v2.0.2
Session info
sessioninfo::session_info()
#> ─ Session info ───────────────────────────────────────────────────────────────
#> setting value
#> version R version 4.2.1 (2022-06-23)
#> os macOS Ventura 13.1
#> system aarch64, darwin20
#> ui X11
#> language en
#> collate en_US.UTF-8
#> ctype en_US.UTF-8
#> tz Europe/Berlin
#> date 2023-09-06
#> pandoc 3.1.1 @ /Applications/RStudio.app/Contents/Resources/app/quarto/bin/tools/ (via rmarkdown)
#>
#> ─ Packages ───────────────────────────────────────────────────────────────────
#> package * version date (UTC) lib source
#> cachem 1.0.8 2023-05-01 [1] CRAN (R 4.2.0)
#> chromote 0.1.2 2023-08-11 [1] CRAN (R 4.2.0)
#> cli 3.6.1 2023-03-23 [1] CRAN (R 4.2.0)
#> colorspace 2.1-0 2023-01-23 [1] CRAN (R 4.2.0)
#> curl 5.0.2 2023-08-14 [1] CRAN (R 4.2.0)
#> data.table 1.14.8 2023-02-17 [1] CRAN (R 4.2.0)
#> digest 0.6.33 2023-07-07 [1] CRAN (R 4.2.0)
#> dplyr 1.1.3 2023-09-03 [1] CRAN (R 4.2.0)
#> evaluate 0.21 2023-05-05 [1] CRAN (R 4.2.0)
#> fansi 1.0.4 2023-01-22 [1] CRAN (R 4.2.0)
#> fastmap 1.1.1 2023-02-24 [1] CRAN (R 4.2.0)
#> fs 1.6.3 2023-07-20 [1] CRAN (R 4.2.0)
#> generics 0.1.3 2022-07-05 [1] CRAN (R 4.2.0)
#> ggplot2 3.4.3 2023-08-14 [1] CRAN (R 4.2.0)
#> glue 1.6.2 2022-02-24 [1] CRAN (R 4.2.0)
#> gt 0.9.0 2023-03-31 [1] CRAN (R 4.2.0)
#> gtable 0.3.4 2023-08-21 [1] CRAN (R 4.2.0)
#> highr 0.10 2022-12-22 [1] CRAN (R 4.2.0)
#> htmltools 0.5.6 2023-08-10 [1] CRAN (R 4.2.0)
#> jsonlite 1.8.7 2023-06-29 [1] CRAN (R 4.2.0)
#> knitr 1.43 2023-05-25 [1] CRAN (R 4.2.0)
#> later 1.3.1 2023-05-02 [1] CRAN (R 4.2.0)
#> lifecycle 1.0.3 2022-10-07 [1] CRAN (R 4.2.0)
#> magick 2.7.5 2023-08-07 [1] CRAN (R 4.2.0)
#> magrittr 2.0.3 2022-03-30 [1] CRAN (R 4.2.0)
#> memoise 2.0.1 2021-11-26 [1] CRAN (R 4.2.0)
#> munsell 0.5.0 2018-06-12 [1] CRAN (R 4.2.0)
#> nflplotR 1.1.0.9006 2023-08-24 [1] Github (nflverse/nflplotR@72bb579)
#> nflreadr 1.3.2.11 2023-09-04 [1] https://nflverse.r-universe.dev (R 4.2.3)
#> pillar 1.9.0 2023-03-22 [1] CRAN (R 4.2.0)
#> pkgconfig 2.0.3 2019-09-22 [1] CRAN (R 4.2.0)
#> processx 3.8.2 2023-06-30 [1] CRAN (R 4.2.0)
#> promises 1.2.1 2023-08-10 [1] CRAN (R 4.2.0)
#> ps 1.7.5 2023-04-18 [1] CRAN (R 4.2.0)
#> purrr 1.0.2 2023-08-10 [1] CRAN (R 4.2.0)
#> R.cache 0.16.0 2022-07-21 [1] CRAN (R 4.2.0)
#> R.methodsS3 1.8.2 2022-06-13 [1] CRAN (R 4.2.0)
#> R.oo 1.25.0 2022-06-12 [1] CRAN (R 4.2.0)
#> R.utils 2.12.2 2022-11-11 [1] CRAN (R 4.2.0)
#> R6 2.5.1 2021-08-19 [1] CRAN (R 4.2.0)
#> Rcpp 1.0.11 2023-07-06 [1] CRAN (R 4.2.0)
#> reprex 2.0.2 2022-08-17 [1] CRAN (R 4.2.0)
#> rlang 1.1.1 2023-04-28 [1] CRAN (R 4.2.0)
#> rmarkdown 2.24 2023-08-14 [1] CRAN (R 4.2.0)
#> rstudioapi 0.15.0 2023-07-07 [1] CRAN (R 4.2.0)
#> sass 0.4.7 2023-07-15 [1] CRAN (R 4.2.0)
#> scales 1.2.1 2022-08-20 [1] CRAN (R 4.2.0)
#> sessioninfo 1.2.2 2021-12-06 [1] CRAN (R 4.2.0)
#> styler 1.10.2 2023-08-29 [1] CRAN (R 4.2.0)
#> tibble 3.2.1 2023-03-20 [1] CRAN (R 4.2.0)
#> tidyselect 1.2.0.9000 2022-11-02 [1] Github (tidyverse/tidyselect@b449033)
#> utf8 1.2.3 2023-01-31 [1] CRAN (R 4.2.0)
#> vctrs 0.6.3 2023-06-14 [1] CRAN (R 4.2.0)
#> webshot2 0.1.1 2023-08-11 [1] CRAN (R 4.2.0)
#> websocket 1.4.1 2021-08-18 [1] CRAN (R 4.2.0)
#> withr 2.5.0 2022-03-03 [1] CRAN (R 4.2.0)
#> xfun 0.40 2023-08-09 [1] CRAN (R 4.2.0)
#> xml2 1.3.5 2023-07-06 [1] CRAN (R 4.2.0)
#> yaml 2.3.7 2023-01-23 [1] CRAN (R 4.2.0)
#>
#> [1] /Library/Frameworks/R.framework/Versions/4.2-arm64/Resources/library
#>
#> ──────────────────────────────────────────────────────────────────────────────Expected result
I would expect to get consistent output in all locations when applying the "correct" function.